Network topology from circuit motifs
Michael Barkasi
2026-04-19
tutorial_network_topology.RmdIntroduction
The simplest mathematical models of neural networks are built from homogeneous McCulloch-Pitts neurons and scalar-weight connections between them. Biological neural networks, such as those in the mammalian cortex, are far more complex. They contain different types of neurons each with their own electrical behavior and spatially extended connections.
Computational neuroscientists standardly model biological neural networks as dynamical systems. The spatial growth-transform (SGT) models used in this package, based on the GT models of Gangopadhyay and Chakrabartty, are one such example. Instead of a one-dimensional weight between two homogeneous neurons, SGT models use a transconductance parameter, a temporal modulation factor, and a transmission velocity parameter to determine network behavior. Roughly, the transconductance parameter is the inverse of a traditional weight, while the temporal modulation factor allows for capturing the electrodynamics of different neuron types at a single spatial point and the transmission velocity parameter allows for modeling the spatial propagation of signals between neurons.
SGT models thus allow for network topologies that not only capture connection strengths between neurons, but also the different types of neurons and their electrodynamics over time and space. A separate tutorial explains SGT models in more detail. This tutorial demonstrates how to build network topologies for these models from circuit motifs.
Initialize network
Let’s set up the R environment by clearing the workspace, setting a random-number generator seed, and loading the DACx package.
# Clear the R workspace to start fresh
rm(list = ls())
# Set seed for reproducibility
set.seed(12345)
# Load DACx package
library(DACx, quietly = TRUE) Network topologies are held in a special class of objects, network, from the DACx package. A new network object can be created with the new.network function.
cortex <- new.network()The network object class is native to C++ and integrated into DACx (an R package) via Rcpp. The object initialized by new.network is a minimal – and empty – “single node” network. In this context, “node” does not mean a single neuron, but rather a cluster of nearby neurons with local recurrent connections. The current structure of our network can be got by calling the fetch.network.components function.
ntw <- fetch.network.components(cortex)## Summary of network: not_provided
## Number of neurons: 0
## Layer names:
## Number of layers: 1
## Number of columns: 1
## Number of patches: 1
## Cell types used:
## Motifs used:The returned object ntw is a list containing all components of the network. For now, the accompanying summary printout is sufficient. As can be seen, networks initialize with zero neurons and no structure. This structure is added in two steps: first set the network node structure with the set.network.structure function, then add meso-scale connections via circuit motifs with the apply.circuit.motif function.
For this tutorial, we will loosely follow the node model of Park and Geffen (2020), which includes three distinct neuron types: excitatory principal pyramidal neurons and two inhibitory types, parvalbumin (PV) and somatostatin (SST) interneurons. These nodes are expected to be (approximately) fully connected, with cells of each type synapsing into cells of all other types.
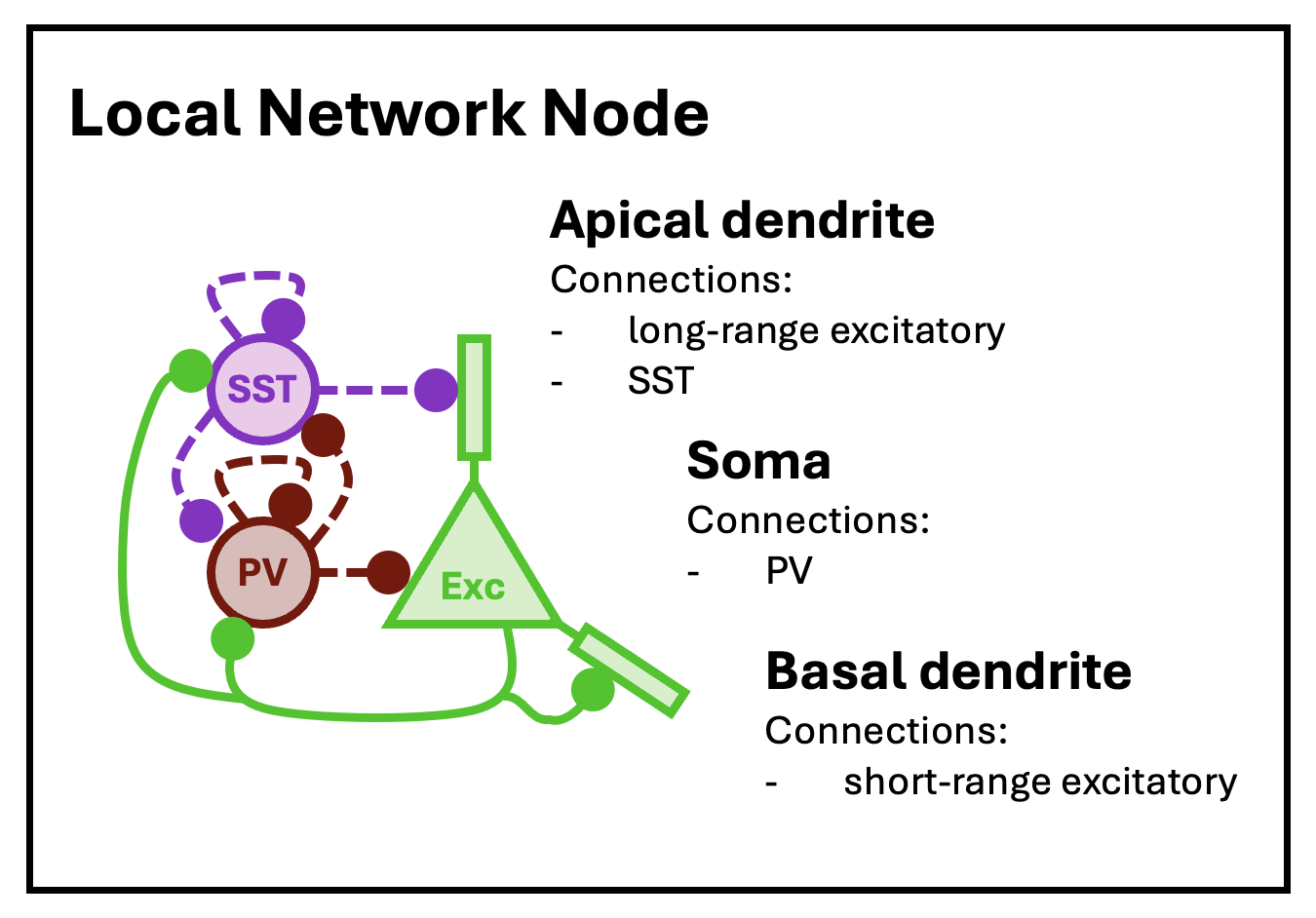
The basic structure of a local network node, as modeled by Park and Geffen (2020).
Nodes are defined by their constitutive cell types and node size (expected number of neurons per type) and arrayed into layers and columns, mimicking the structure of the cortex. With a network initialized, this structure is set with the set.network.structure function. To capture the topology of the Park and Geffen node model, we will create nodes with three cell types (principal, PV, and SST), with expected counts of 10, 5, and 5, respectively, except for layer 1 (L1), in which expected counts will be 5, 0, and 1. To mimic the major functional areas of the cortex, we create six layers and array columns in two dimensions:
cortex <- set.network.structure(
cortex,
neuron_types = c("principal", "PV", "SST"),
layer_names = c("L6", "L5", "L4", "L3", "L2", "L1"),
n_columns = 8,
patch_depth = 4,
layer_separation_factor = 3.0,
column_separation_factor = 5.5,
patch_separation_factor = 5.5,
neurons_per_node = matrix(c(
10, 5, 5, # L6
10, 5, 5, # L5
10, 5, 5, # L4
10, 5, 5, # L3
10, 5, 5, # L2
5, 0, 1 # L1
), ncol = 3, byrow = TRUE)
)Notice that we set both a value for n_columns and a value for patch_depth. Each variable controls the size of an independent columnar axis perpendicular to the laminar axis. By convention, if only one of the two columnar axes has functional significance (e.g., preserved retinotopy or preserved tonotopy), it’s assumed to be the one controled by n_columns.
The DACx package pre-defines a number of different cell types, including principal cells, PV, and SST interneurons. The process of loading and setting cell types is discussed in the tutorial on SGT models. The principal class is special, insofar as it’s not a cell type itself, but instead points towards the most numerous (i.e., the “principal”) cell type in each layer. Known principal-layer combinations can be seen by calling the principal.neurons function:
principal.neurons(print_nicely = TRUE)## Principal neuron types by layer:
## layer: pyramidal
## L1: Neurogliaform_cell
## L2: pyramidal
## L3: pyramidal
## L4: spiny_stellate
## L5: pyramidal
## L6: pyramidal_L6If we check the summary for our network again, we see that its cell types include the principal cell types for each layer:
ntw <- fetch.network.components(cortex)## Summary of network: not_provided
## Number of neurons: 3468
## Layer names: L6, L5, L4, L3, L2, L1
## Number of layers: 6
## Number of columns: 8
## Number of patches: 4
## Cell types used: pyramidal_L6, PV, SST, pyramidal, spiny_stellate, Neurogliaform_cell
## Motifs used: local connectionsWe also see that the “Motifs used” field now contains “local connections”. All cells initialize with a local arbor structure which includes at least one axon and one dendrite spatially constrained to more-or-less the space of the cell’s node. The exact shape and branch structure of these arbors are defined by the cell type, but in all cases are built by a biased random walk. This walk generates a tree of nodes and “local connections” are formed by connecting axon nodes to nearby dendrite nodes to form synapses. These connections mostly stay local to a cell’s node, with two exceptions. Nearby cells in neighboring nodes stand a good chance of synapsing, and pyramedial cells are initialized with long apical dendrites which span multiple layers and thus can synapse with cells in other nodes (or neighboring nodes) within the same column.
At this point, it may be helpful to visualize a sampling of the network we have created. We will start with a 2D visualization which collapses the patch_depth dimension, but later we will look at the network in 3D.
plt <- plot.network(cortex)
plt$plot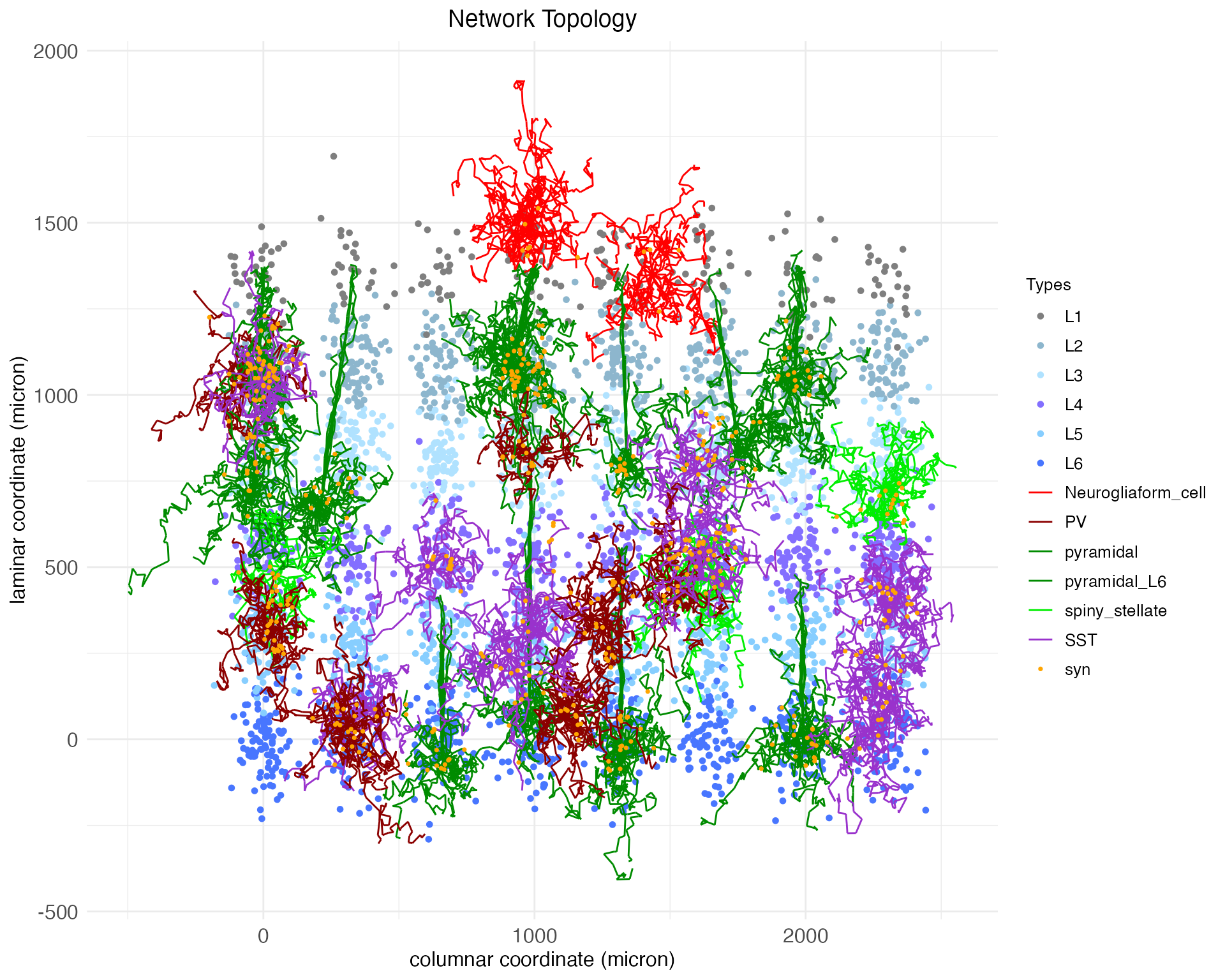
By default, the plot.network function plots all cell bodies as dots, colored by layer, and plots the arbors for a random 1% of those cell bodies, colored by cell type. Synapses (syn) are plotted as dots. The synapses shown are determined by the presynaptic axons that are plotted. Notice that the arbors shown are mostly small and compact, expect for the pyramidal neurons, which have long apical dendrates which extend upwards.
Notice that the plot gives coordinates in microns. Spatial coordinate giving a cell’s soma (i.e., body) location along the columnar and laminar axis are continuous and real-valued. These coordinates are used in conjunction soma-to-synapse distance integrated along the axon and the transmission velocity parameter to simulate spike propagation. Five parameters control the coordinates assigned to each soma: column_diameter c_{\mathrm{diameter}}, layer_height l_{\mathrm{height}}, column_separation_factor F_{\mathrm{column}}, layer_separation_factor F_{\mathrm{layer}}, and patch_separation_factor F_{\mathrm{patch}}. A node is created for each combination of patch integer-index p, layer integer-index l, and column integer-index c. Each node is assigned a coordinate \langle x_c,y_l\rangle: z_p = p \frac{c_{\mathrm{diameter}}F_{\mathrm{patch}}}{2} y_l = l \frac{l_{\mathrm{height}}F_{\mathrm{layer}}}{2} x_c = c \frac{c_{\mathrm{diameter}}F_{\mathrm{column}}}{2} Within each node, a coordinate \langle z_n,y_n,x_n\rangle is assigned to each neuron n by sampling from a normal distribution: z_n \sim \mathcal{N}(z_p, \frac{c_{\mathrm{diameter}}}{2}) y_n \sim \mathcal{N}(y_l, \frac{l_{\mathrm{height}}}{2}) x_n \sim \mathcal{N}(x_c, \frac{c_{\mathrm{diameter}}}{2}) By default, DACx sets c_{\mathrm{diameter}} = 120.0, l_{\mathrm{height}} = 180.0, F_{\mathrm{patch}} = 2.5, F_{\mathrm{column}} = 2.5, and F_{\mathrm{layer}} = 2.5. The above network cortex used larger values for the column and patch separation factors so that the columns would be more visually distinct in plots.
These default values were obtained via back-of-the-envelope calculations from widefield microscopy images of wild-type mouse auditory cortex. They can be extended to estimate values needed to simulate a complete brain area. By eyeballing those same images (multiplying pixels by a resolution of 0.4 microns per pixel), we estimate a mean neuronal soma diameter of 14 microns, cluster radius of 62 microns, and cluster density (i.e., proportion of cluster occupied by neuronal somas) of 0.4. These numbers imply approximately 278 cells per cluster. Assuming a span of 13 columns across the tonotopic axis of the auditory cortex (approximately 2 mm) and a patch depth of 3 perpendicular to it (approximately 0.5 mm), six columns at that cluster density is approximately 65,000 neurons total, which is a fair approximation for a single hemisphere of mouse auditory cortex.
Of course, for demonstration purposes we will stick to the much smaller network we’ve already created, but these calculations show how the parameters of the set.network.structure function can be used to scale up to a more realistic network size. Returning to our demonstration network, we have only “local” connections between neighboring neurons. Next, we will add meso-scale connections between layers and columns using circuit motifs.
Define and load motifs
A circuit motif, represented in the DACx package by special objects of class motif, is a predefined pattern of connections between layers and columns in a network. It is a recipe for directing axons – and thereby connecting cells – between nodes, based on cell type, and pre and post-synaptic node coordinates (patch, layer, column indices).
Below we will define and load three canonical circuit motifs thought to characterize sensory cortex: (1) a translaminar and intracolumnar projection motif of principal (excitatory) columnar projections, (2) a lateral projection motif for cross-columnar communication, and (3) a translaminar deep-layer PV arborization motif for intracolumnar inhibition. These motifs are summarized in the figure below.
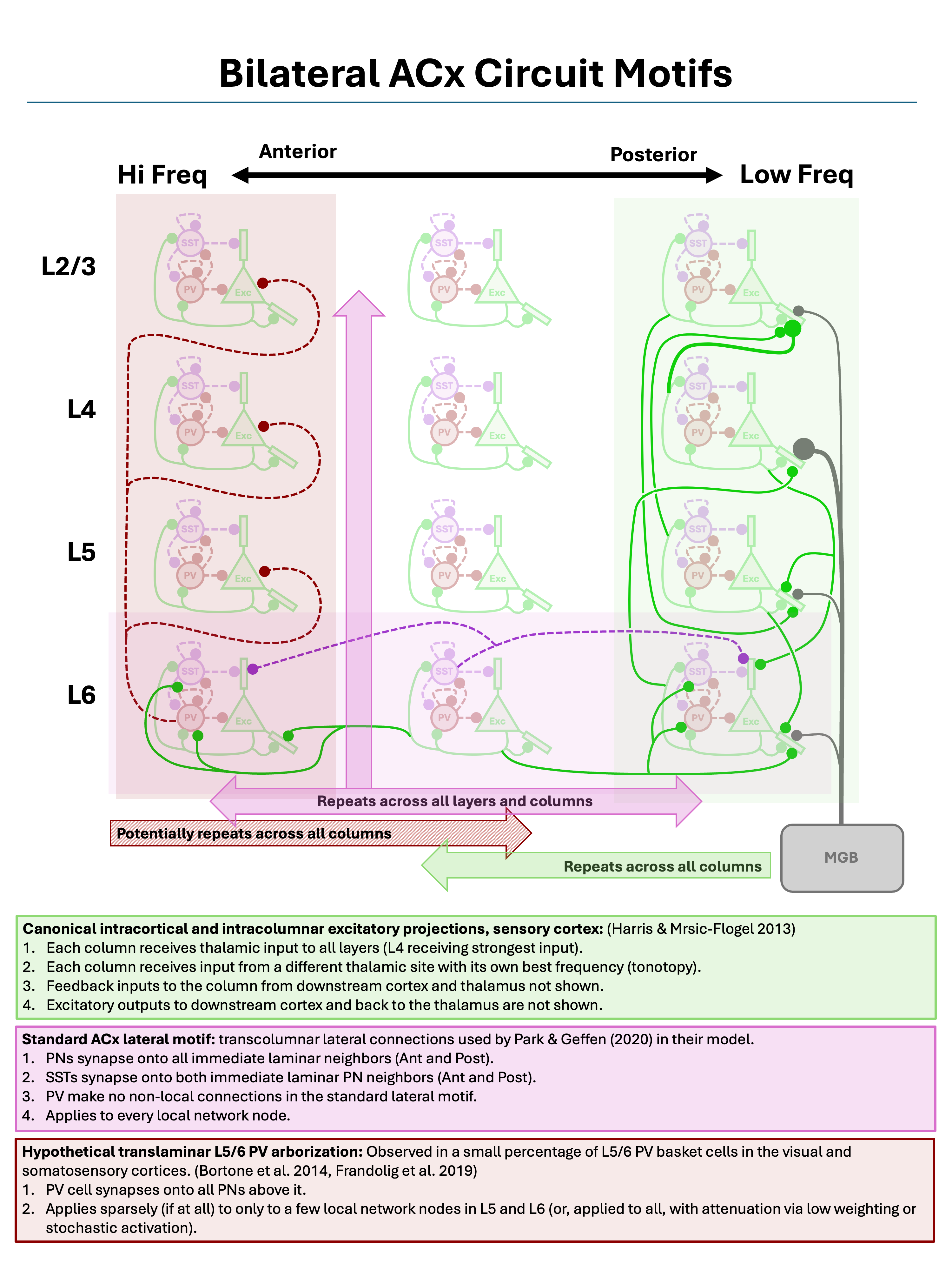
Three canonical sensory cortex circuit motifs, as described by Harris and Mrsic-Flogel (2013), Park and Geffen (2020), Bortone et al. (2014), and Frandolig et al. (2019).
As shown by the figure, these motifs are “columnar”, in the sense that they are defined relative to a single cortical column and repeated across all columns, both along the primary columnar axis sized by n_columns and the secondary columnar axis sized by patch_depth.
Principal columnar projections
The new.motif function initializes an empty motif object. Let’s initialize one to model the canonical translaminar and intracolumnar principal (excitatory) projections in the sensory cortex. These connections are largely what defines a “column” in the cortex, and thus they connect neurons in different layers while staying within the same column.
motif.pp <- new.motif(motif_name = "principal projections")Connections between layers can then be added to the motif using the load.projection.into.motif function. This function takes as arguments the motif object, the source layer name, and the target layer. By default, it extends the axons of principal cells in the source layer to the target layer between those layers with a connection strength of 0.5. The connection strength value, held in the variable connection_strength, is used to control the expected initial strength of synapses along the extended axon. This value can be modified via a fourth argument to the function. Here, for example, we load projections with connection strength 0.9 from layer L4 to layer L2 and layer L3 into our new motif:
motif.pp <- load.projection.into.motif(motif.pp, "L4", "L2", 0.9)
motif.pp <- load.projection.into.motif(motif.pp, "L4", "L3", 0.9)We have increased the strength of this connection from the default because layer 4 (the primary site of input from outside the cortex into the cortex) has its strongest intra-cortical projections to layers 2 and 3. By default, the loaded projections are between principal neurons, which for layer 4 are spiny stellate cells and for layers 2 and 3 are pyramidal neurons. Below we will show how the pre and post-synaptic cell types can be modified through additional function arguments. For now, let’s load weaker projections from layer 4 into the deep layers below it:
motif.pp <- load.projection.into.motif(motif.pp, "L4", "L5", 0.25)
motif.pp <- load.projection.into.motif(motif.pp, "L4", "L6", 0.25)The column-defining principle projections also typically include recurrent loops between layers 2 and 3 and layer 5, which we’ll assume are of average (i.e., default) strength:
motif.pp <- load.projection.into.motif(motif.pp, "L2", "L5")
motif.pp <- load.projection.into.motif(motif.pp, "L3", "L5")
motif.pp <- load.projection.into.motif(motif.pp, "L5", "L2")
motif.pp <- load.projection.into.motif(motif.pp, "L5", "L3")Finally, these projections also include a feed-forward projection from layer 5 to layer 6, and a feedback projection from layer 6 to layer 4:
motif.pp <- load.projection.into.motif(motif.pp, "L5", "L6", 0.25)
motif.pp <- load.projection.into.motif(motif.pp, "L6", "L4", 0.25)With the complete principle-projection motif defined, we apply it using the apply.circuit.motif function. This function takes as arguments the network object and the motif object. It adds the connections defined in the motif to the network.
cortex <- apply.circuit.motif(
cortex,
motif.pp
)Let’s update our original plot of the network. The plot.network function returns not only the generated plot, but also the indexes for the plotted somas and arbors. This allows for plots to be reproduced as a network is built out, showing the same cell arbors as more motifs are added.
plt <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
)
plt$plot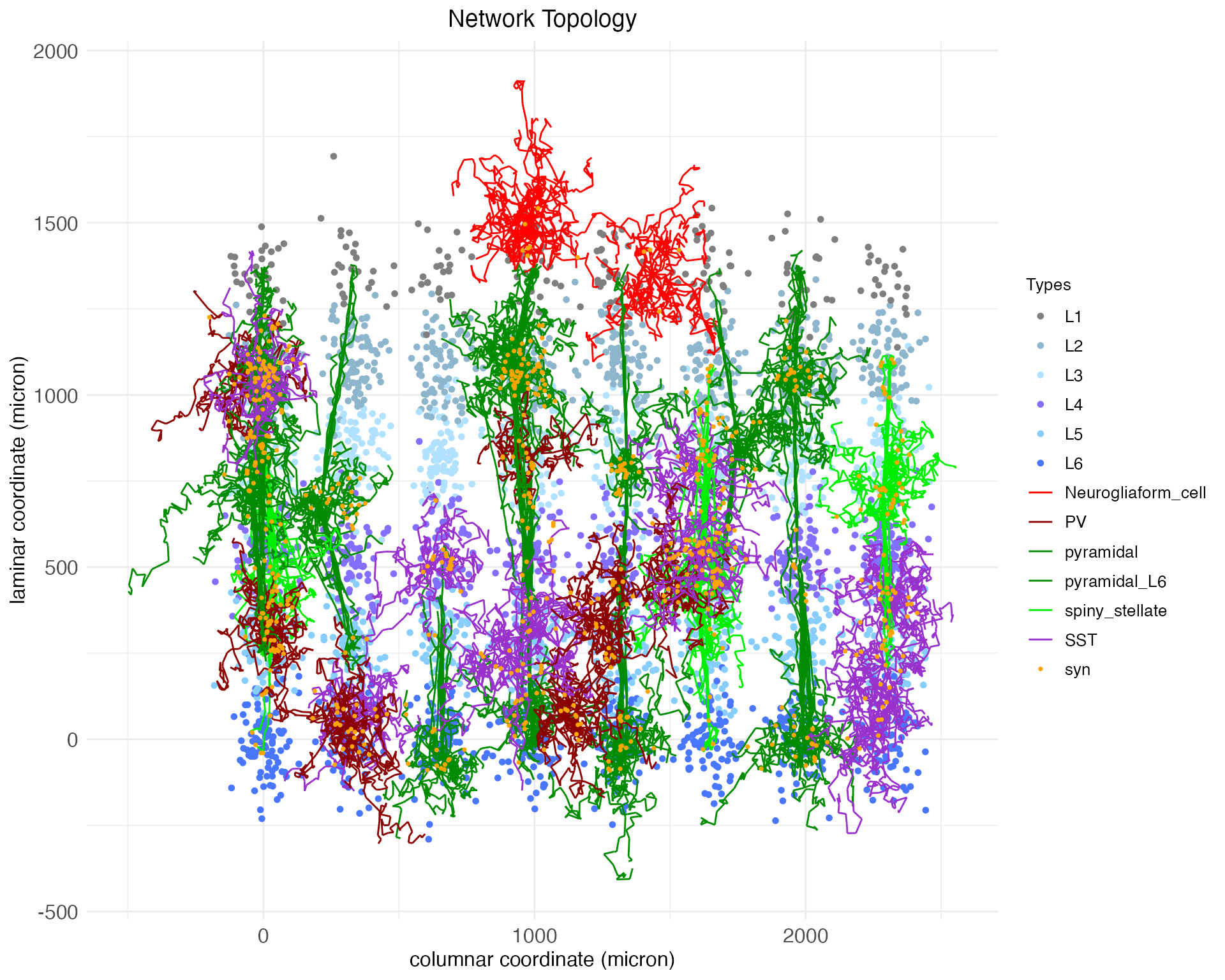
We can also plot the network (with this sampling of arbors) in 3D, showing the full spatial shape of the arbors:
plt <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
threedim = TRUE
)
plt$plotNotice, as is clear in both the 2D and 3D plots, that while the arbors of some cells cross between layers, they all mostly remain in column. There are no meso-scale lateral projections between columns.
Auditory lateral projections
Within sensory cortex, cortical columns tend to respond to a single type of stimulus. For example, cortical columns in the visual cortex might have preferred edge orientations while columns in the auditory cortex prefer certain sound frequencies. More advanced cortical computations require integrating input from a range of stimuli (e.g., a range of edge orientations or sound frequencies), and thus there must be some mechanism of column cross-talk. This is thought to be accomplished by lateral connections between cells of the same layer, but different columns. For example, Park and Geffen (2020) propose that these lateral connections include excitatory projections from principle neurons out to cells of all types in adjacent columns and inhibitory projections from SST interneurons out specifically to principle (excitatory) neurons in adjacent columns. Thus, for this motif, we need to make use of the presynaptic_type and postsynaptic_type arguments of the load.projection.into.motif function to specify the relevant neuron types for each projection.
By default, load.projection.into.motif creates projections within the same column. To create lateral projections, we need to use the max_col_shift_up and max_col_shift_down arguments of the function to specify how many columns away from the source column the target column can be. For the purposes of this demonstration, we’ll set these to four. However, lateral projections are assumed to drop off quickly, and so when the apply.circuit.motif function is called, the actual length of any lateral branch projection will be a random value placing its expected endpoint somewhere between the source column and four columns away. Actually, this scaling applies with all arbor-branch generation, including the translaminar, intracolumnar projects of the previous motif.
motif.ACxlat <- new.motif(motif_name = "ACx laterals")
# Add projection for each layer
for (layer in c("L1", "L2", "L3", "L4", "L5", "L6")) {
# Excitatory laterals
for (celltype in c("principal", "PV", "SST")) {
motif.ACxlat <- load.projection.into.motif(
motif.ACxlat,
presynaptic_layer = layer,
postsynaptic_layer = layer,
presynaptic_type = "principal",
postsynaptic_type = celltype,
max_col_shift_up = 4,
max_col_shift_down = 4
)
}
# Inhibitory laterals
motif.ACxlat <- load.projection.into.motif(
motif.ACxlat,
presynaptic_layer = layer,
postsynaptic_layer = layer,
presynaptic_type = "SST",
postsynaptic_type = "principal",
max_col_shift_up = 4,
max_col_shift_down = 4
)
}As before, we can apply this motif and visualize the results:
cortex <- apply.circuit.motif(
cortex,
motif.ACxlat
)
plt <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
threedim = TRUE
)
plt$plotNow we have not only vertical projections between layers, but horizontal projections between columns.
Intracolumnar inhibitory projections
Cortical computations require careful balance between excitatory and inhibitory activity, both within local nodes and between nodes within columns. Balancing the principal columnar projections – which are predominantly excitatory – are hypothesized to be intracolumnar inhibitory projections from PV interneurons out of the deep layers below L4 (Bortone et al. 2014, Frandolig et al. 2019). These function to shut the column down. Let’s define a motif for these inhibitory projections. For this motif, we again need to make use of the presynaptic_type and postsynaptic_type arguments of the load.projection.into.motif function to specify that this projection motif is from inhibitory PV neurons to excitatory principal neurons.
motif.dinhib <- new.motif(motif_name = "deep inhibition")
# Layer 5 projections
motif.dinhib <- load.projection.into.motif(
motif.dinhib,
presynaptic_layer = "L5",
postsynaptic_layer = c("L4", "L3", "L2"),
presynaptic_type = "PV",
postsynaptic_type = "principal"
)
# Layer 6 projections
motif.dinhib <- load.projection.into.motif(
motif.dinhib,
presynaptic_layer = "L6",
postsynaptic_layer = c("L5", "L4", "L3", "L2"),
presynaptic_type = "PV",
postsynaptic_type = "principal"
)As before, we can apply this motif and visualize the results.
cortex <- apply.circuit.motif(
cortex,
motif.dinhib
)
plt <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
threedim = TRUE
)
plt$plotAs can be seen, now the three deep-layer PV cell arbors (in dark red) are extending their upwards towards the superficial layers, as expected from the motif.
Notice that in this case we’ve loaded multiple projections at once by specifying a vector of target layers. While the argument presynaptic_layer can take only a single value, if postsynaptic_layer is given a vector of layer names, projections will be loaded from the presynaptic layer to each of the specified postsynaptic layers. In this case, the pre and post-synaptic neuron types will be the same for each projection, so if different types are required, then multiple calls to the load.projection.into.motif function will be necessary.
Network topology summary
Let’s return again to the fetch.network.components function and look at the network now:
ntw <- fetch.network.components(cortex, include_arbors = TRUE)## Summary of network: not_provided
## Number of neurons: 3468
## Number of synapses: 86359
## Layer names: L6, L5, L4, L3, L2, L1
## Number of layers: 6
## Number of columns: 8
## Number of patches: 4
## Cell types used: pyramidal_L6, PV, SST, pyramidal, spiny_stellate, Neurogliaform_cell
## Motifs used: local connections, principal projections, ACx laterals, deep inhibitionIn addition to what’s printed, the function also extracts other important summary information. For example, here are the mean numbers of synapses per pre-synaptic neuron, per cell type:
## neuron_type n_synapses
## 1 Neurogliaform_cell 5.407186
## 2 PV 22.029797
## 3 pyramidal 30.286422
## 4 pyramidal_L6 21.802589
## 5 spiny_stellate 51.580645
## 6 SST 16.411271Before concluding, two other plotting options are helpful in exploring a network’s topology. First, for both 2D and 3D plots, the complex arbors can be replaced by straight-line edges between pre and post-synaptic cells, allowing synaptic connections to be seen directly. Further, these connections can be restricted by motif, e.g., here we see only synaptic connections added as a result of applying the lateral projection motif:
plt_lat <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
plot_motif = "ACx laterals",
reconstruct_arbors = FALSE
)
plt_lat$plot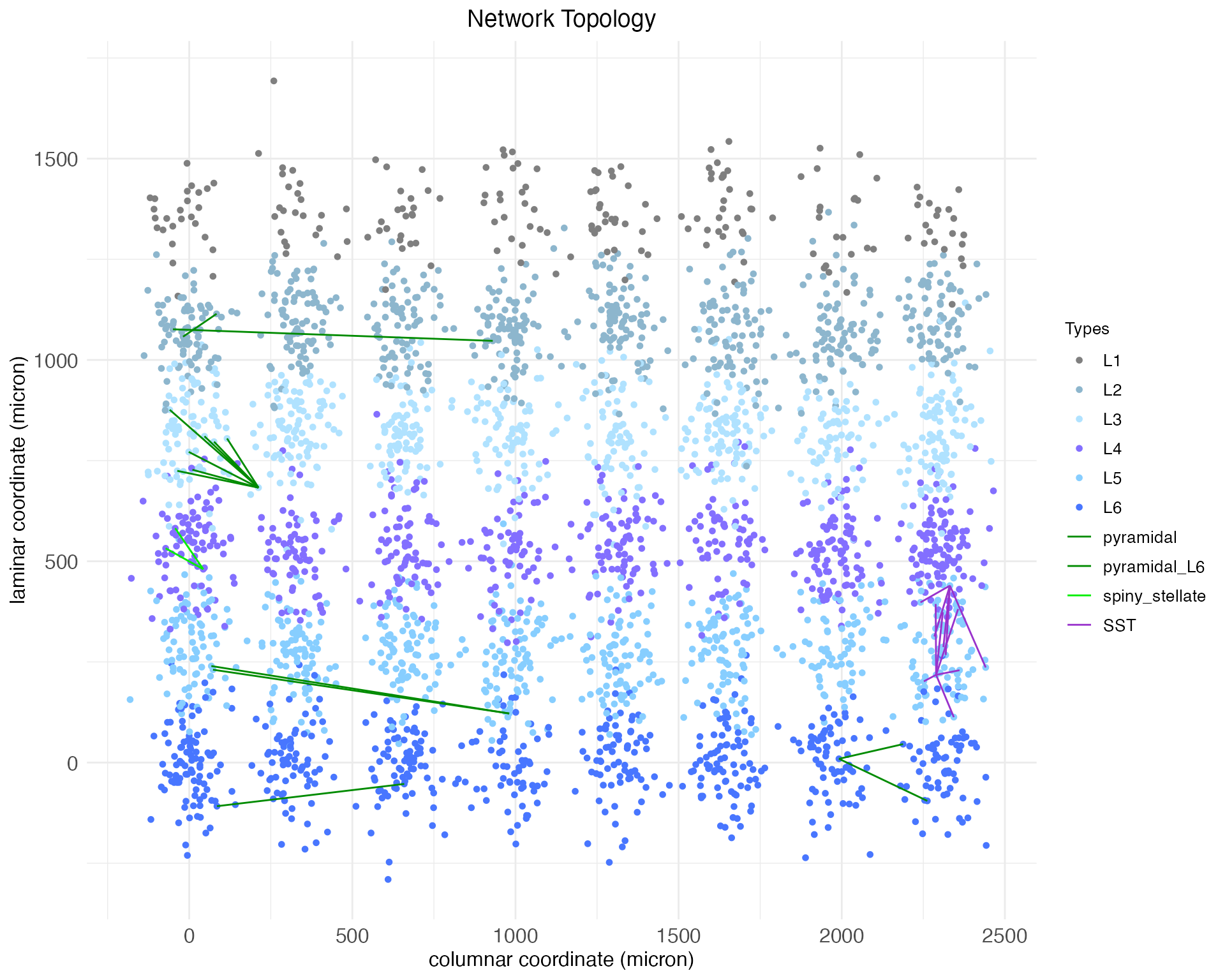
As can be seen, although the arboring induced by the motif was extensive, the number of new synaptic connections was not. This suggests that we should either extend the maximum column shift for this motif, or decrease the proximity needed between an axon and dendrite for a synapse to form. The latter can be done using the synaptic_neighborhood argument of the set.network.structure function.
Second, for both 2D and 3D plots, cell arbors can be colored by process type, i.e., whether they are an axon or dendrite. To conclude, here is our network ploted in 3D with arbors colored by process type:
plt <- plot.network(
cortex,
soma_mask = plt$soma_mask,
arbor_idx = plt$arbor_idx,
threedim = TRUE,
edge_color = "is_axon"
)
plt$plot